Examples¶
This page walks through fitting real ALMA data with galfit-uv using two models: a single free-n Sersic and a Sersic(n=1) + point source decomposition.
Target¶
GQC J0054-4955 observed in CO(3-2) at z ~ 2.5.
Demo data: The measurement set can be downloaded from: https://drive.google.com/file/d/1TPZQvP-7wc5Kk169Gh6IgiYAUoaug2bb/view?usp=sharing
- 1980 visibility points after time-averaging
- UV range: 5 -- 111 kilo-lambda
- Wavelength: 2.817 mm
- Zero-baseline flux: ~19.2 mJy
Two models are compared:
| Model | Components | Free params |
|---|---|---|
| Model A | Single Sersic (free n) | 7 |
| Model B | Sersic(n=1) + Point source | 7 (n fixed) |
Step 1: Export Visibilities¶
Extract averaged visibility data from the measurement set.
from galfit_uv.export import export_vis, save_uvtable
dvis, wle = export_vis('data/GQC_J0054-4955_avg.ms', verbose=True)
print(f"Extracted {len(dvis.u)} visibility points")
print(f"Wavelength: {wle*1e3:.3f} mm")
print(f"uv range: {dvis.uvdist.min()/1e3:.0f} -- {dvis.uvdist.max()/1e3:.0f} klambda")
print(f"Total flux (zero-baseline): {abs(dvis.vis[0])*1e3:.2f} mJy")
# Optionally save a uvplot-compatible table
save_uvtable(dvis.u, dvis.v, dvis.vis, dvis.wgt, 'uv_table.txt', wle=wle)
Output:
Extracted 1980 visibility points
Wavelength: 2.817 mm
uv range: 5 -- 111 klambda
Total flux (zero-baseline): 19.17 mJy
Step 2: Build the Model¶
Model A — Single Sersic:
from galfit_uv.models import make_model_fn
model_fn, param_info = make_model_fn(['sersic'])
# labels = ['flux', 'Re', 'n', 'incl', 'PA', 'dx', 'dy']
Model B — Sersic(n=1) + Point:
from galfit_uv.models import make_model_fn
model_fn, param_info = make_model_fn(
['sersic', 'point'],
fixed={'sersic:n': 1.0},
)
# labels = ['sersic:flux', 'sersic:Re', 'point:flux', 'incl', 'PA', 'dx', 'dy']
Step 3: Run MCMC¶
from galfit_uv.fit import fit_mcmc
result = fit_mcmc(
dvis, model_fn, param_info,
max_steps=5000,
burnin=2500,
nwalk_factor=5,
outpath='./fit_output',
seed=42,
)
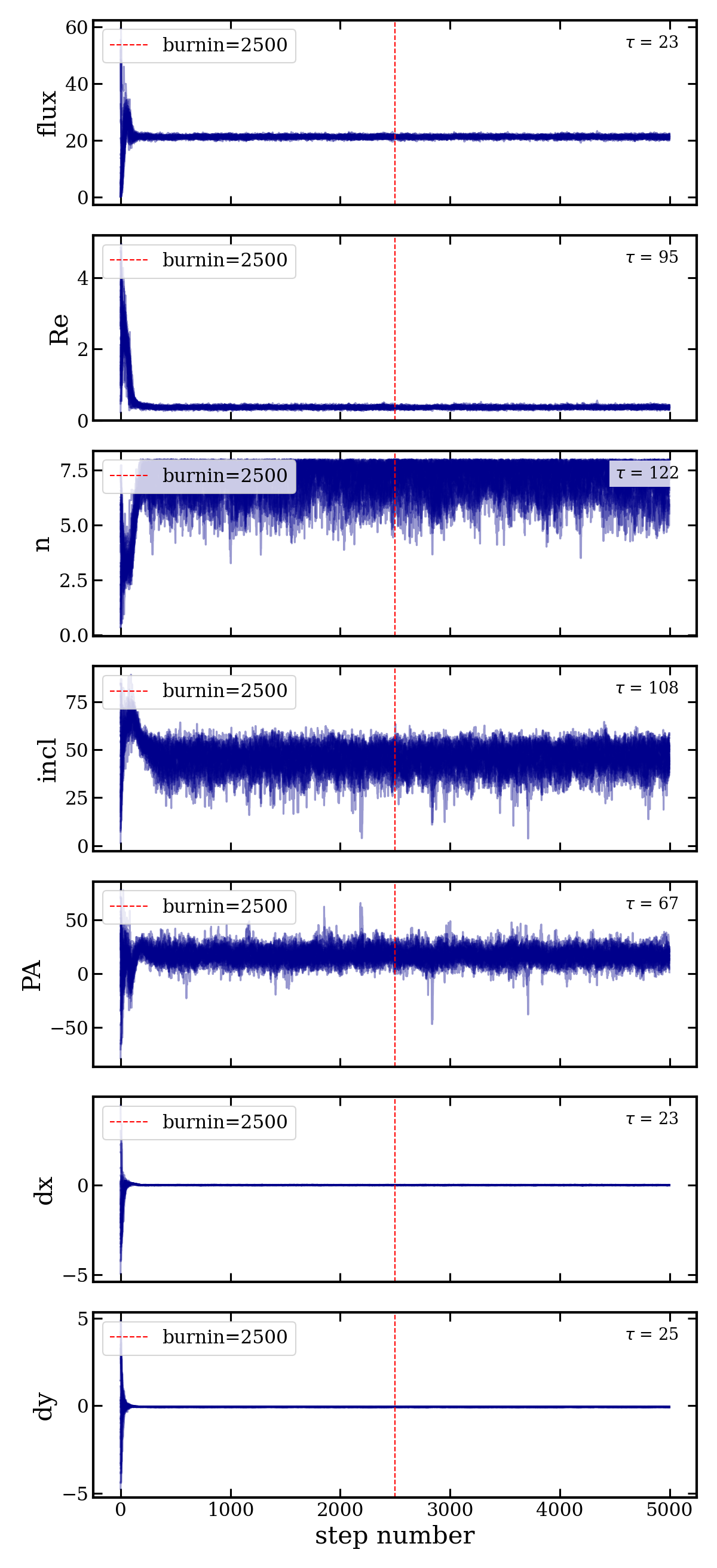
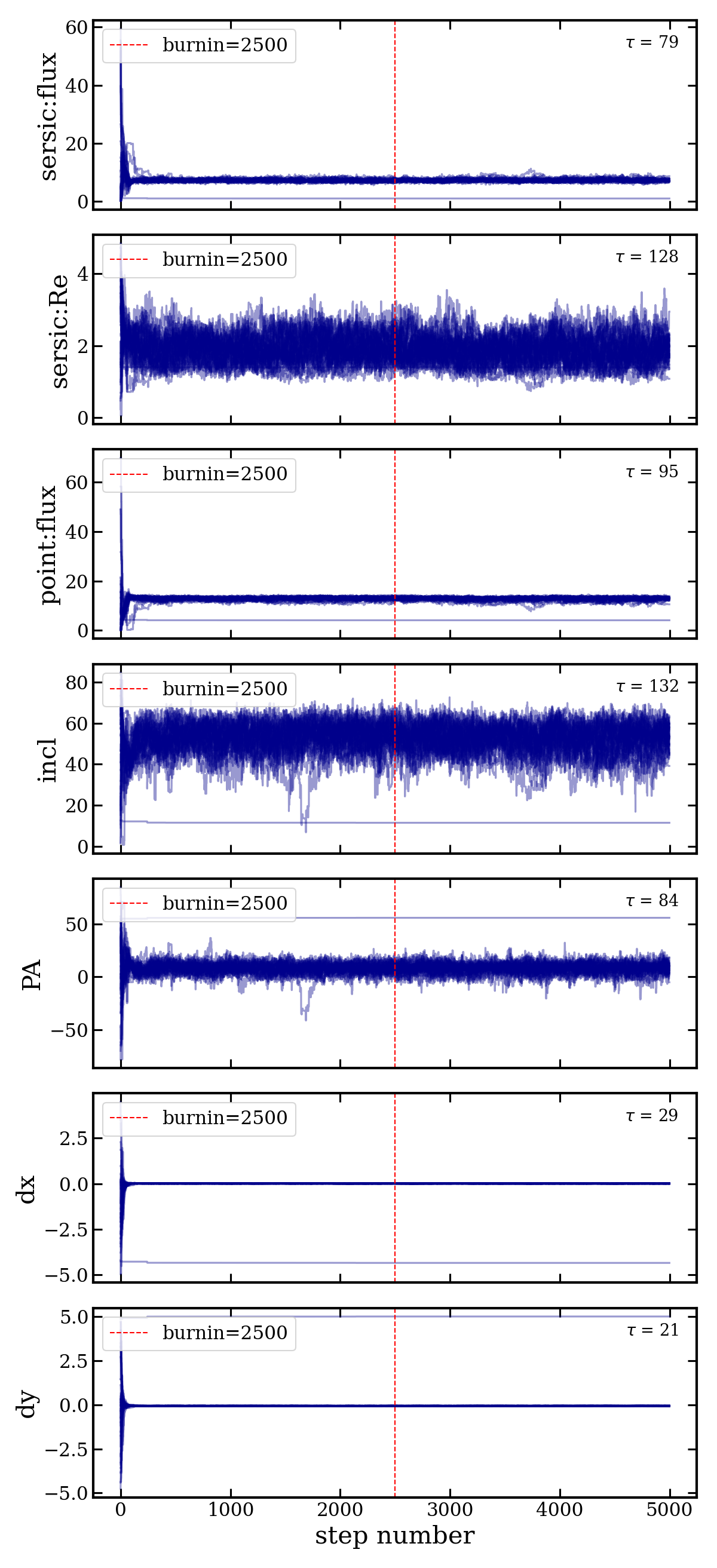
Settings: 35 walkers (7 free params x 5), 5000 steps per walker, 2500 burn-in steps.
Outputs saved to outpath/:
fit_results.fits— multi-extension FITS with data, model, samples, fit statisticschains.png— walker traces with autocorrelation time annotationscorner_plot.png— posterior corner plot
Model A: Single Sersic (free n)¶
Results¶
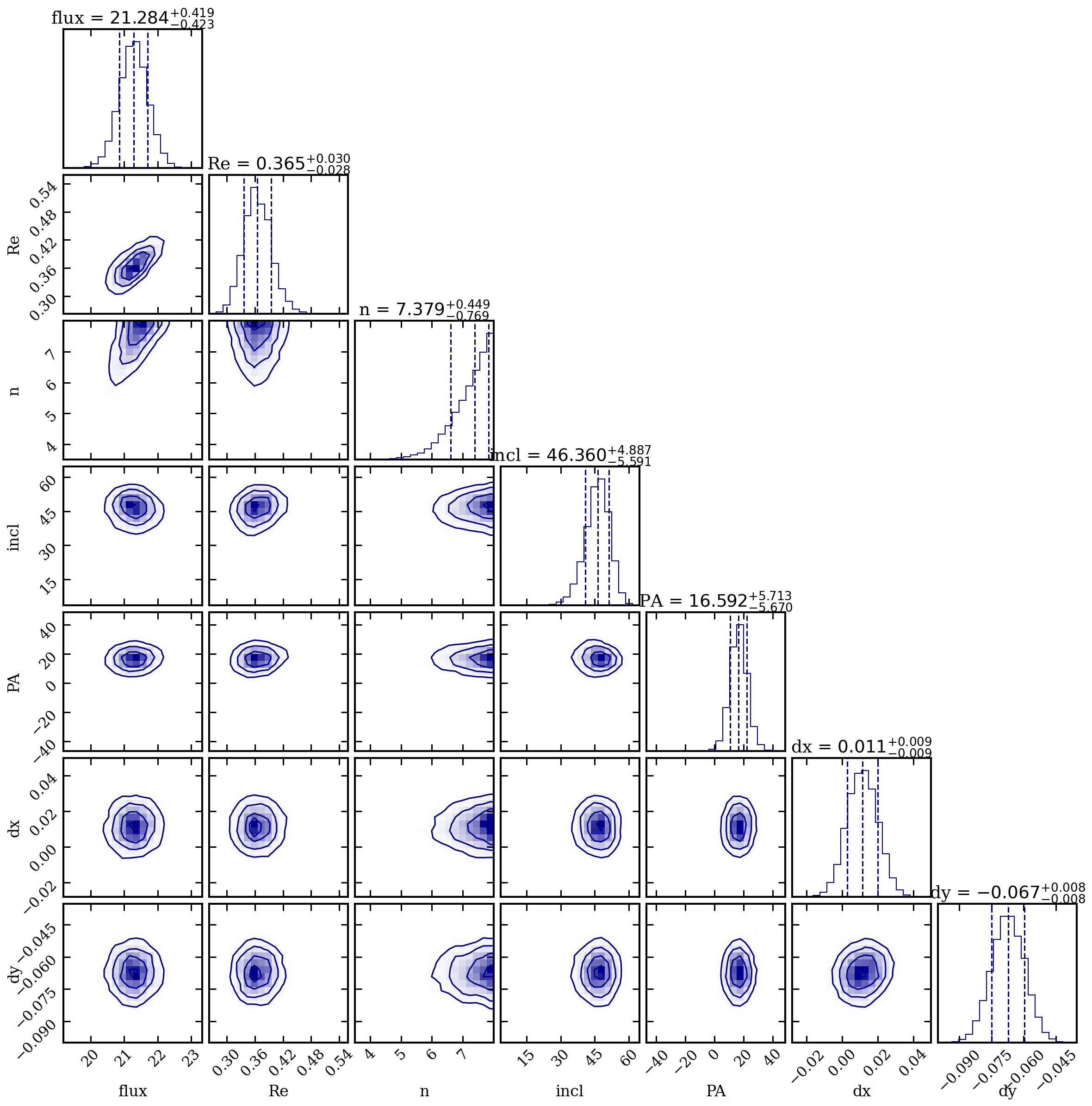
| Parameter | Best-fit | +1sig | -1sig |
|---|---|---|---|
| flux (mJy) | 21.28 | +0.42 | -0.42 |
| Re (arcsec) | 0.365 | +0.03 | -0.03 |
| n | 7.38 | +0.45 | -0.77 |
| incl (deg) | 46.4 | +4.9 | -5.6 |
| PA (deg) | 16.6 | +5.7 | -5.7 |
| dx (arcsec) | 0.011 | +0.009 | -0.009 |
| dy (arcsec) | -0.067 | +0.008 | -0.008 |
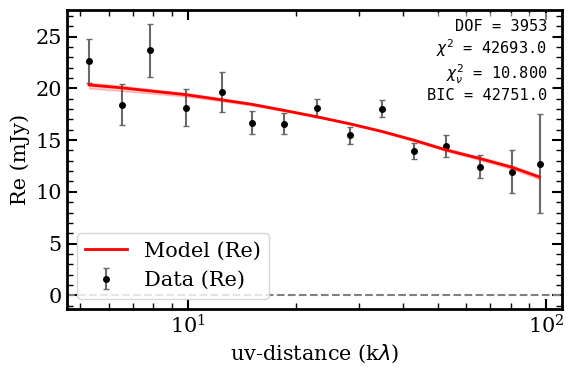
Fit statistics: chi2/DOF = 10.80, BIC = 42751
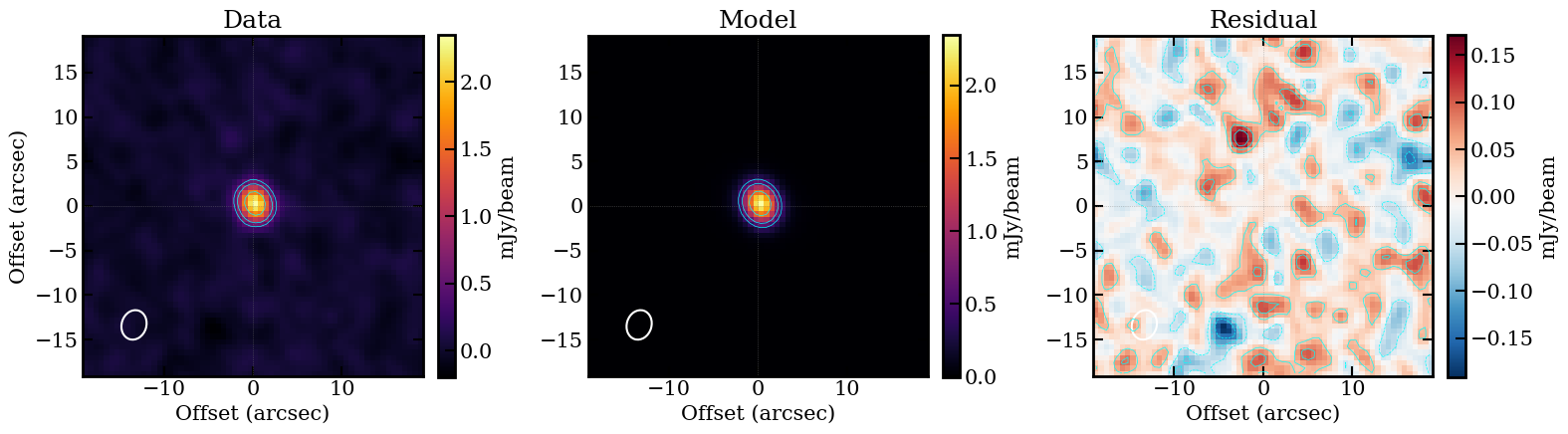
The high Sersic index (n ~ 7.4, well above de Vaucouleurs n=4) indicates a very concentrated core with extended wings, while the small effective radius (Re ~ 0.37 arcsec) is consistent with a compact emission region.
Figures¶
Model B: Sersic(n=1) + Point Source¶
Results¶
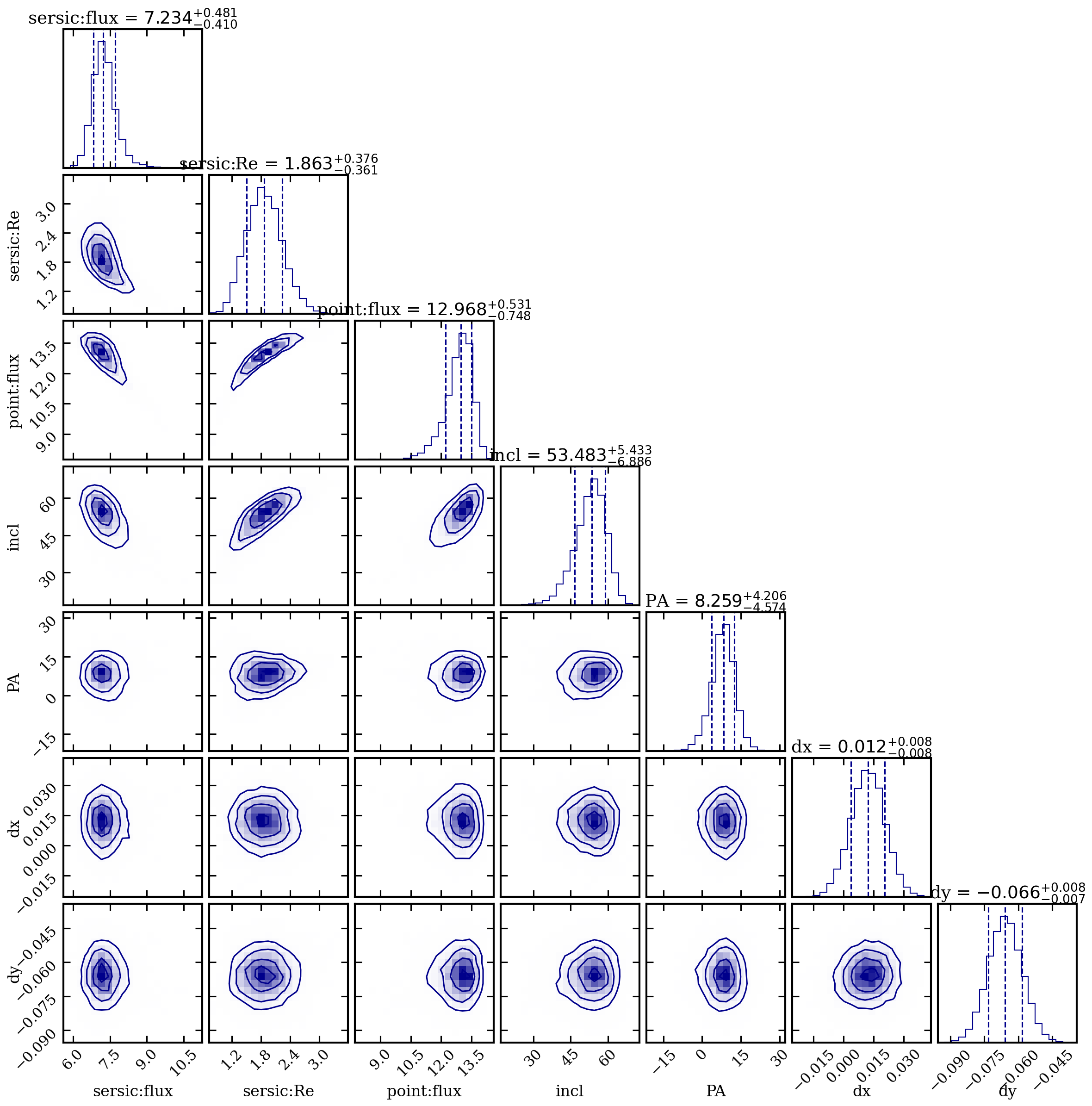
| Parameter | Best-fit | +1sig | -1sig |
|---|---|---|---|
| sersic:flux (mJy) | 7.23 | +0.48 | -0.41 |
| sersic:Re (arcsec) | 1.86 | +0.38 | -0.36 |
| sersic:n | 1.00 | — | — (fixed) |
| point:flux (mJy) | 12.97 | +0.53 | -0.75 |
| incl (deg) | 53.5 | +5.4 | -6.9 |
| PA (deg) | 8.3 | +4.2 | -4.6 |
| dx (arcsec) | 0.012 | +0.008 | -0.009 |
| dy (arcsec) | -0.066 | +0.008 | -0.007 |
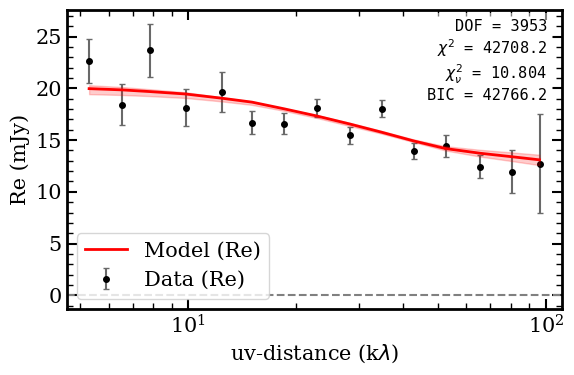
Fit statistics: chi2/DOF = 10.80, BIC = 42766
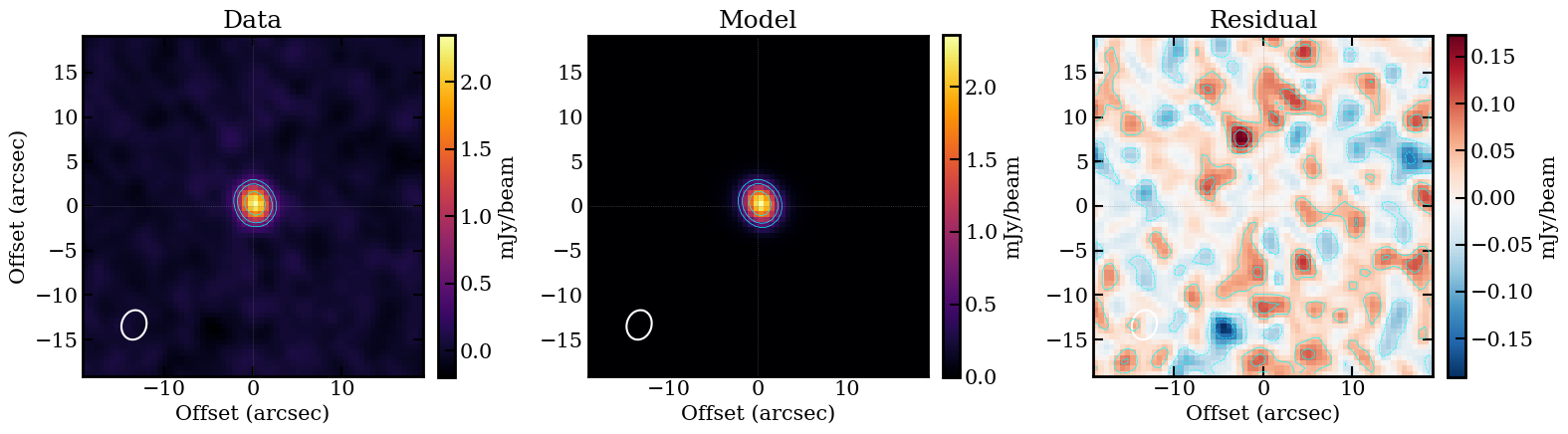
The model decomposes the source into an extended exponential disk (~7.2 mJy, Re ~ 1.86 arcsec) plus a dominant unresolved point source (~13 mJy), with a fitted inclination of ~53 degrees.
Figures¶
Model Comparison¶
| Metric | Model A (Sersic) | Model B (Sersic+Point) |
|---|---|---|
| Total flux (mJy) | 21.3 | 7.2 + 13.0 = 20.2 |
| Extended size | Re = 0.37 arcsec | Re = 1.86 arcsec |
| Sersic index | n = 7.4 (free) | n = 1.0 (fixed) |
| BIC | 42751 | 42766 |
Both models yield nearly identical reduced chi-squared values. The single Sersic has a slightly lower BIC, but the two-component decomposition provides a more physically interpretable separation between an extended disk and a compact nucleus. The geometry parameters (center offset) are consistent between the two models.